VASISTHA viz The Archaeogenetics Blog ::
Steppe-->Brahmin? Final evidence against Harvard's claim
I received a reply from Harvard's Dr. David Reich to the mail I had sent (check previous post comments).
Hi Ashish,
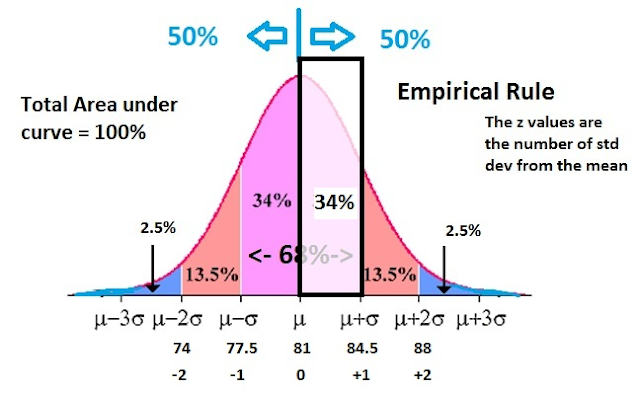
The Z-scores are formal tests for model fit – appropriately calibrated by a block jackknife – and the tail of large Z-scores are simply reflecting the reality of model failure.
What the analyses are showing is that the “Indian Cline” is in fact best modeled as a two-dimensional cloud not a one-dimensional gradient: a mixture of three source populations that cannot be simplified to a mixture of two source populations as we had done in previous modeling for example in Reich et al. Nature 2019.
As such, what our enrichment analyses are showing is simply that the groups of traditionally priestly status tend to be enriched for the Steppe-related source population after controlling for the proportions from the other two.
Yours, David
I will address these points in this analysis post. This will be final evidence to disprove the confident Steppe--->Brahmin causality claim made in the study.
Let us make a new model to study excess steppe ancestry based on the qpAdm ancestries of the 140 modern populations studied in Narasimhan et al 2019 [1](
the study). For the purpose of this analysis, we will ignore the bad qpAdm p-values in the study and assume that the models and ancestry estimates provided are accurate for all 140 populations, although some are clearly not.
We will assume that a 2 source ANI ASI model is accurate for modeling all modern South Asians.
ANI = a IVC + (1-a) Steppe
ASI = b AHG+ (1-b) IVC
where
ANI is Ancestral North Indian
ASI is Ancestral South Asian
a is ancestry coefficient of IVC ancestry (or Indus Periphery) in ANI
b is ancestry coefficient of AHG in ASI
0<=a,b<=1
Our main aim is to use the qpAdm coefficients for each population, and by keeping the AHG and IVC ancestries the same as the qpAdm model (thereby fixing the amount of steppe, which will be different than the qpAdm model) but varying a,b in such a fashion that the overall errors in the 140 groups are minimized.
let G be one of the 140 modern groups.
G = e.ANI + f.ASI; 0<=e,f<=1 & e+f=1
G = ea.IVCN + eb.Steppe + fc.IVCS + fd.AHG
ea + fc = qpAdm IVC estimate for group G
fd = qpAdm AHG estimate for group G
IVCN = IVC ancestry type from ANI part
IVCS =IVC ancestry type from ASI part
For easier understanding, let me explain it with kalash.
qpAdm for Kalash: 0.123 AHG + .599IVC + 0.278 Steppe
we can write it as
where (1-b)/b is the ratio of IVC to AHG ancestry in ASI, and (1-a)/a is the ratio of Steppe/IVC ancestry in ANI.
If both these ratios are 0.4, it would mean
ASI = 71.24% AHG + 28.76%IVC
and ANI = 71.24%IVC + 28.76% Steppe
and Kalash would become
0.123 AHG + 0.0492 IVCS + 0.5498 IVCN + 0.2199 Steppe.
We can see that the steppe component of 0.2199 is less than the qpAdm estimate of 0.278. ie the model predicts that the ANI for Kalash had a higher steppe component than what we chose. This difference of 0.0581 is the fit error for this group for this particular choice of ratios.
Fitting all 140 Groups with the ANI-ASI model
For a particular set of ratios (1-a)/a and (1-b)/b, we will get a set of 140 fit errors for 140 different groups. We can now vary both the ratios and come to a final choice of ratios at which the sum squared of the 140 fit errors is minimized. We also place the constraint that none of the 4 ancestry coefficients can be negative.
This can be done via Excel VBA macro, or more simply by using the solver plugin in excel.
The error minimizing algorithm gives the result for IVC/AHG ratio in ASI as 0.421 and that of Steppe/IVC in ANI as 0.372. ie ANI = 72.9% IVC + 27.1% Steppe and ASI = 70.36% AHG and 29.64%.
The excess steppe error ranges from -0.13 to 0.18 steppe % points with a mean of 0.0015 and a median of -0.0037 and a standard deviation of 0.0527. mean and std dev estimated from 5000 Monte Carlo random simulation runs of picking errors of 15 random groups each time.
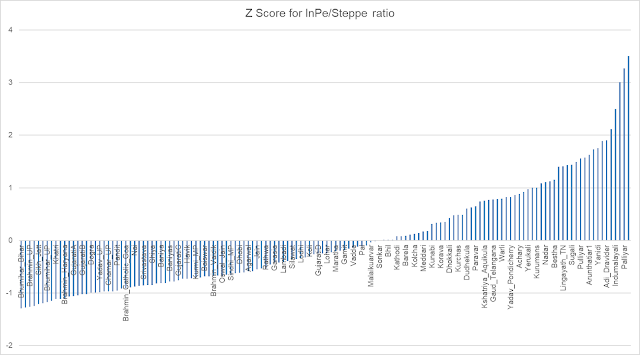
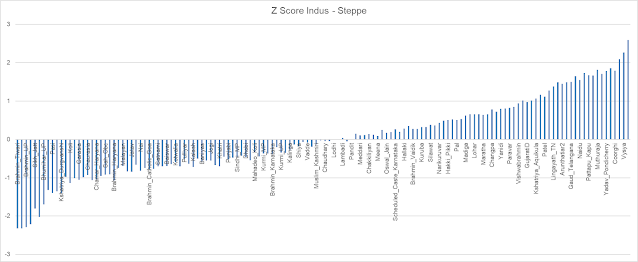
Converting these error scores to Z-Scores by standardizing, results in a Z-Score range of -2.41 to 2.18, with mean 0 and std dev 1 - confirming that it is indeed Standard Normal Z distribution.
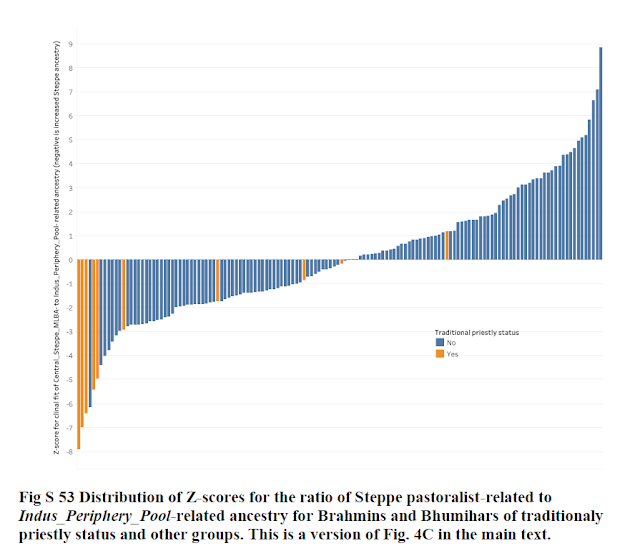
Whereas the Z-Score range for the study in question is -7.9 to 8.2, with a mean 0 but std dev 2.79. The Z-Scores are inflated for some reason, so they no longer represent a deviation from the mean of the 140 groups.
I think what has happened here is that the Z-Scores given in the study are Z-scores for deviation from the 2 way ANI ASI model, calculated as (Excess steppe)/(Std Error of the steppe qpAdm estimate).
Or in other words
Z = (ANI ASI Model estimate - qpAdm estimate)/(SE of the qpAdm estimate)
The mean of 0 is just a coincidence, helped by the fact that the underlying excess steppe calculated also has a mean for the140 groups close to 0.
This is what was confirmed to me by Dr. David Reich in his email reply - ie. the large Z Scores indicate failure of the model.
However, this raises serious concerns -
The deviation for a group from the fit of a FAILED MODEL should not be used to make any definitive conclusions. The conclusions made based on the Z scores of these failed fits are wrong. The excess steppe signal being picked up is a mirage. I will prove my point in the subsequent section by showing a WORKING 3 Source model.
A Working 3 Source model for 140 modern Indian groups
Dr. Reich confirmed in his email reply that the ANI ASI model does not work for modern Indians, and no 2 source model will work as the groups lie inside a cloud rather than a cline. A minimum 3 source model is required to model these 140 groups without outliers. I agree with this assessment.
Similar to above my model, I will now formulate the methodology for a 3 source model. Bear in mind that there can be multiple solutions to this problem, I am outlining just 1 or 2.
I define the 3 populations as
ASI = IVC + AHG in a fixed ratio of ancestries, with AHG dominating the component
AI =IVC + AHG in another fixed ratio of ancestries, with IVC dominating the component
ANI = IVC + Steppe, in a fixed ratio of ancestries.
We fix the sum of the AHG components to the qpAdm estimate for the group,
the sum of IVC components to qpAdm estimate of IVC for the group,
Steppe in the fit model will be a result of the remaining IVC component multiplied by the steppe/IVC ratio for ANI.
The floating variables used will be the 3 ratios, as well as 140 multipliers which will be specific to 1 group respectively and denote the % of the total AHG ancestry utilized by ASI.
eg.
qpAdm estimates for Kalash: 0.123 AHG + 0.599IVC + 0.278 Steppe
can be broken into
[g * 0.123 AHG + g * 0.123 * r1 IVC] +
ASI
[(0.123 - g * 0.123)AHG + 0.123 x (1-g) * r2 IVC] +
AI
[{0.599 - (g * 0.123 * r1) - (0.123*(1-g).r2)} IVC +{0.599-(g*r1) -(0.123x(1-g)*r2)}*r3 Steppe]
ANI
where g is the proportion of AHG for Kalash which is part of ASI, 0<g<1. there are 140 of these - g1 to g140.
r1 = ratio of IVC to AHG in ASI; common for all 140 groups
r2 = ratio of IVC to AHG in AI; common for all 140 groups
r3 = ratio of Steppe to IVC in ANI; common to all 140 groups
The coefficient we get for Steppe is compared to the qpAdm estimate and the difference is the error. Our error minimization algorithm will vary these 143 parameters such that the sum squared of the 140 errors is minimized.
We use the Excel Solver plugin for this. One additional constraint is that none of the 6 coefficients should be negative.
RESULT
After the error minimization, we get a working model.
The mean of the error is 0.0001, the median is 0.0000, the standard deviation is 0.0014. The range of the error is from -0.0014 to max 0.017. The Z-Scores for model fit (error/SE) range from -0.0969 to 1.30, giving proof of successful fits for all 140 populations.
The error range of -0.13 to 0.18 in the best fitting ANI ASI model is reduced to -0.0014 to 0.017, and the standard deviation is reduced from >0.05 to 0.0014.
This is expected because we are already assuming that a 3 source model given by qpAdm (AHG + IVC + Steppe) is accepted, therefore any 3-way combination of these 3 underlying ancestries will also give a successful fit. However, the 3 sources I have chosen have a chance of providing more historical information as a proximal model.
From the output of the fitted model we get:
ANI = 64% IVC + 36% Steppe
ASI = 99% AHG + 1% IVC
AI = 61% IVC + 39% AHG
and all 140 groups as a combination of these 3 ancestries.
The PCA of all 140 groups with these 3 proximal sources is below
The good fit of the model is confirmed by all 140 groups lying inside the triangle in the PCA. It is immediately evident that the 4 Brahmin groups based on which the study made its claims about excess Steppe ancestry in Brahmins have a unique position on this PCA.
The other outlier in the study's Z-Score, was Panta_Kapu with a Z-Score of 8.85, showing the largest deviation among 140 groups from the ANI ASI model.
However, in the 3 source model, Panta Kapu is seen on the other cline - the ANI AI cline, with no contribution from ASI. It showed as an opposite end outlier in the study because of its low ASI, whereas the 4 Brahmin groups were outliers because of the high need for ASI.
THE BRAHMIN OUTLIER GROUPS
It has to be said that it is questionable whether these 4 particular groups are representative of Brahmins in the UP and Bihar regions. There are many UP brahmins on Anthrogenica who claim that these samples are erroneous and their own ancestry does not match these samples at all. On Eurogenes G25 too, the difference between the UP Brahmins from
Mondal et al 2017 data differs from the other UP brahmins. Maybe there is a difference between eastern and western UP brahmin ancestry, but I have not studied this matter and therefore have no judgment. With this caveat, let's proceed.
As per the 3 source model above, none of the 4 outliers - Bhumihar_Bihar, Brahmin_UP, Brahmin_Tiwari, and Brahmin_Nepal, are in need of excess steppe ancestry over and above what is provided by ANI. In other words; Nepal, UP, Chattisgarh & Bihar Brahmins DO NOT NEED A SEPARATE ANI SOURCE WITH HIGHER STEPPE/IVC ANCESTRY RATIO.
The excess Steppe needed in the failed model was actually a false signal, what is needed in these 4 outlier groups is an ancestry source that has excess ASI(AHG) and almost nil IVC on top of ANI as well. This is exactly the point that I had made in my Part 1 critique, that eastern Indians have excess AHG ancestry as well, which is driving this false signal of excess steppe requirement in the failed ANI ASI model.
If it were not for this high AHG source requirement specific to these 4 groups, they would not have needed a high Steppe/IVC ANI source to model them.
An important corollary is this: If a set of populations can only be modeled as a 3-source admixture, then only 1 ancestry (or its ratio with any 2nd) cannot be the only explanation for a 2-source model failure. By mathematical certainty, at least 2 sources / 2 ratios will explain the deviation from the 2 source model.
How can it be? How can this ANI group meet with almost pure original AHG (also called AASI) ancestry? It appears that eastern India still has populations with the highest AHG like ancestry even in modern times, even more so than the southern Dravidian Paniya & Irula tribals. The Birhor tribals of Jharkhand, Bihar, Chattisgarh are Austroasiatic-speaking hunter-gatherers. They possess even greater AASI/AHG ancestry than Paniya, mixed with SE Asian ancestry, and there is some evidence to suggest that even the little IVC like ancestry in Birhor is of recent admixture (DATES: S1 Shahr_Sokhta_BA2, S2 ONG.SG, Target BIR.SG: Output 23.77 +- 8.65 generations, 1092CE - 1576CE)
This is additional evidence to the almost consensus hypothesis that AASI entered India from the East/Southeast of India in paleolithic times.
This can explain how these 4 Brahmin groups from east India picked up AASI/Austroasiatic ancestry. At some point, they must have intermarried locals from the region that they settled in. These Brahmin groups are just showing the adaptation to their region, just like other groups show their adaptation to their specific regions. Another line of evidence to prove this hypothesis is that eastern Indo Aryan languages show Munda language substrate which is absent from western IA languages, hinting at IA speaking groups changing the language of the eastern Indian region where Munda people lived.[2][3][4]
qpAdm for Birhor: P-value 0.0003 (best fit i got): 0.69 Onge + 0.175 IndusPWest + 0.138 Dai
PCA Based on Language
Based on the Language association with these 140 groups taken from the study, I plotted the PCA of the groups and classified them according to language. ANI component is CORRELATED with IE speakers, however, it of course does not imply causation. It could just be a matter of geography that North India has a higher ANI component (steppe can enter only through north) as well as IE speaking. ANI also has a 65% IVC component, the major component.
Think of it like this, if we assume that the hypothesis that North India was already IE speaking by 2500BCE before the ANI component existed is true, then it would also be true that the later ANI component entry would cause a higher ancestry % in the North. So regardless of whether north India spoke IE language pre steppe or not, the ANI component will always be higher in the north of India. Implying correlation = causation lacks logical rigour.
Then is there a way to check if this ANI ancestry has any special relation to Language and religion?
There could be. I agree with the study's assertion that Priestly groups were the "
traditional
custodians of liturgy in Sanskrit". Rather than look at ratios, a simpler way is to just check the overall % of ANI ancestry in a group and correlate it with various measures. The idea is simple, a group with 75% ANI ancestry can be hypothesized to have a significantly higher chance of being of Priestly status than a group with 35% ANI, and this hypothesis can then be checked.
Let's test if Priestly groups have significant excess ANI ancestry over the rest of the groups of the Indian subcontinent. We do this via Welch's two-tailed T-test to compare the means of ANI ancestry in 2 groups, and statistically check if the means are significantly different at 0.05 significance (95% confidence). If the means are significantly different, the null hypothesis will be rejected and the p-value of the test will be below 0.05.
For Brahmins vs Non-Brahmins, the mean of ANI is significantly different (p<<<0.05).
This proves that the ANI component is maximized in Brahmins and hence it must be causal to the Brahmin groups right? Not quite.
There is a big problem with the null hypothesis that I formulated. The null hypothesis that Brahmins will have on average the same ANI as the rest of the Indian populations spanning all the country is a really bad one, because we know that's not going to be true. The Brahmins derive from the Vedic people who were located in NW India as per the Rig Veda. Since those times, brahmins then moved out to various parts of the country to spread religion and culture. As discussed above, a northern population is for sure going to be heavier in ANI than the aggregate of the country, which includes the South and East to drag the number down for that team. The proof is below.
N/NW India = Groups from Pakistan, Delhi, Haryana, J&K, Punjab, Rajasthan.
So what Null hypothesis should we test?
To test whether the ANI component is significantly related to Brahmins vs others, we should compare the mean of ANI ancestry in Brahmins vs the mean of ANI ancestry in all other groups of N/NW India and Pakistan, including the so-called 'backward groups'. If that null hypothesis of the equal mean between the 2 sets can be rejected, then we can confidently make the claim that indeed ANI is significantly related to Brahmin populations.
p=0.23>>0.05, hence the null hypothesis cannot be rejected. The mean of the ANI ancestry in Brahmins & NW Indians of various caste hierarchies is similar.
Bear in mind that the Brahmin groups don't include lower ANI groups like Marathi, Tamil, Gujarati brahmins, etc, whereas the NW groups don't include higher ANI groups like Ror, Punjab_Jatt, Kamboj, etc. For sure, we could be missing the other side of the same coin too, but I don't think there are any other Brahmin groups with more ANI than UP brahmins. The more data we have the better the statistical power of the tests will be.
CONCLUSION
Nevertheless, the fact that traditional custodians of liturgy in Sanskrit (Brahmins) tend to have more Steppe ancestry than is predicted by a simple ASI-ANI mixture model provides an independent line of evidence—beyond the distinctive ancestry profile shared between South Asia and Bronze Eastern Europe mirroring the shared features of Indo-Iranian and Balto- Slavic languages (58)—for a Bronze Age Steppe origin for South Asia’s Indo-European languages.
In my above work, I have poked various holes in this emphasized part of the claim made in Narasimhan et al 2019.
References
1. Narasimhan VM, Patterson N, Moorjani P, et al. The formation of human populations in South and Central Asia. Science. 2019;365(6457):eaat7487.
doi:10.1126/science.aat7487
2. Peterson, John. "Fitting the pieces together – Towards a linguistic prehistory of eastern-central South Asia (and beyond) "
Journal of South Asian Languages and Linguistics, vol. 4, no. 2, 2017, pp. 211-257.
https://doi.org/10.1515/jsall-2017-0008
3. Ivani, Jessica K., Paudyal, Netra, and Peterson, John. "Indo-Aryan – a house divided? Evidence for the east-west Indo-Aryan divide and its significance for the study of northern South Asia"
Journal of South Asian Languages and Linguistics, vol. 7, no. 2, 2020, pp. 235-274.
https://doi.org/10.1515/jsall-2021-2029
4.
The West/East Divide among Indo-Aryan languages