VASISTHA
R1a L657 in Roopkund lake, Uttarakhand from 800CE, but no Steppe.
In 2019, a paper
1 was published with the analysis of the remains of 38 individuals whose bones were found in Roopkund lake, Uttarakhand at an altitude on 5000m above sea level.
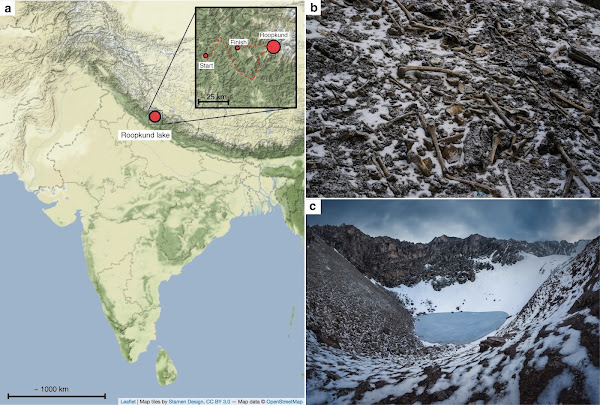 |
| Context of Roopkund Lake. a Map showing the location of Roopkund Lake. The approximate route of the Nanda Devi Raj Jat pilgrimage relative to Roopkund Lake is shown in the inset. b Image of disarticulated skeletal elements scattered around the Roopkund Lake site. Photo by Himadri Sinha Roy. c Image of Roopkund Lake and surrounding mountains. Photo by Atish Waghwase |
13 of these skeletons were confirmed to have died around 1800CE in a single event. These 13 individuals had east mediterranean ancestry, ie. they were foreigners. They were also accompanied by 1 person of Malaysian/vietnamese descent.
On the contrary, it was found that 23 of these Individuals were Indians seemingly from differing jAtis and geographies within India. 10 were women, and 13 were men. They all died between 700CE and 900CE, possibly in separate incidents. It is likely that these people were going on a 'teerth' to nanda Devi temple and had an accident/bad weather during the mountainous trek. It is this group which is called Roopkund_A that I will analyze in this post. So far, it is the only high coverage group of ancient DNA that we have from India.
What makes this group A so fascinating is that all these individuals are unrelated and are like a snapshot of whole of India at that particular time (800CE). The sample size is also decent. The PCA below will explain.
PCA for Roopkund A
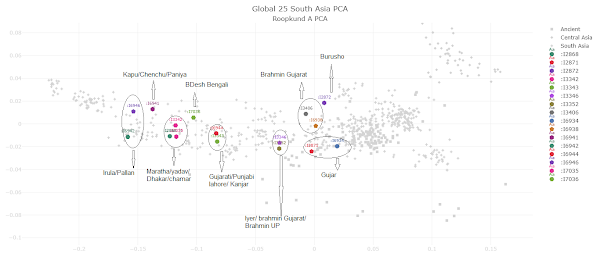 |
| 16 of the 23 A group samples, and note of which modern population is closest to each. Click to enlarge. |
All 23 Roopkund_A samples and best qpAdm models.
Sample ID | Gender | Y HG | p-Value | I8726 | Sintashta | Irula | WSHG | Mongolia_N | Tarim_EMBA |
I3351 | M | J2b2a~ | 0.8420 | 31% | 34% | 34% | | | |
I6938 | F | - | 0.2296 | 27% | 29% | 38% | | | 6% |
I2872 | F | - | 0.0772 | 41% | 28% | 31% | | | |
I3406 | M | J2a1a1b1 | 0.1210 | 47% | 26% | 15% | 4% | 10% | |
I6943 | M | J2b2a2b~ | 0.8163 | 37% | 24% | 39% | | | |
I2871 | F | - | 0.1623 | 44% | 23% | 33% | | | |
I3349 | F | - | 0.3871 | 31% | 22% | 47% | | | |
I3346 | M | E1b1b1b2a1a~ | 0.0375 | 32% | 22% | 47% | | | |
I6945 | F | - | 0.0743 | 10% | 20% | 70% | | | |
I6934 | F | - | 0.4091 | 54% | 18% | 28% | | | |
I6944 | F | - | 0.3330 | 21% | 18% | 61% | | | |
I3352 | M | R2a2b1b2b | 0.1488 | 48% | 15% | 37% | | | |
I7036 | M | R2 | 0.0670 | 18% | 14% | 59% | | 10% | |
I3344 | F | - | 0.0945 | 45% | 11% | 44% | | | |
I3343 | F | - | 0.2011 | 28% | 11% | 61% | | | |
I3402 | M | H3b | 0.0067 | 41% | 11% | 33% | | 15% | |
I6941 | M | R2 | 0.1235 | 8% | 10% | 74% | | 8% | |
I3342 | M | H1a1a4b | 0.1230 | 29% | 9% | 53% | | 10% | |
I7035 | F | - | 0.1260 | 20% | 6% | 74% | | | |
I2868 | M | H1a1b1 | 0.2126 | 20% | 6% | 74% | | | |
I3407 | M | H1a1a4b | 0.0880 | 15% | 4% | 81% | | | |
I6946 | M | R1a1a | 0.2905 | | | 100% | | | |
I6942 | M | R1a1a1b2a1a1a1f~ | 0.1593 | | | 100% | | | |
I8726 = Indus periphery sample from Shahr-Sokhta Iran with least AASI (Onge like) component
Irula are Dravidian (also called Irula language) speaking tribals from Tamil nadu
WSHG = Western Siberian Hunter gatherers, samples found in Tyumen and Sosonivoy (Russia)
Mongolia_N = Mongolia North Neolithic samples with east asian ancestry
Tarim_EMBA = Tarim basin samples from the Xiaohe culture 1900BCE
All qpAdm result files can be found here
Irula itself can be modeled as (Rakhigarhi + Onge) or (Darra-i-kur_afghanistan + Onge) with -ve coefficients for Sintashta, or negligible noise level steppe ancestry considering std errors (results in folder link above)
 On performing statistical analysis on the above data we can say that
On performing statistical analysis on the above data we can say that
- Irula component is statistically significantly associated with Y hg R1a at 99% confidence interval (independent t-test p value = 0.0000107561) compared to non R1a. Click for report.
- Sintashta component is statistically significantly associated with the J2 y HG at 99% confidence interval (independent t-test p value = 0.00595690) compared with non J2. Click for report. This is only possible if Sintashta autosomal ancestry was ultimately incorporated through steppe rich females (as Sintashta male samples are all R1a). A similar result is seen in the Swat valley IA data (Narasimhan et al 2019).
- Y Hgs R2 and H are not statistically significantly associated with any component. Low sample size becomes relevant here as null hypothesis could not be rejected like in above 2 cases.
Above is my proof to all those who say and keep saying that there is a very high correlation between Steppe ancestry and R1a in modern Indians.
The only proper aDna sampleset we have from India shows exactly the opposite correlation.
On Y-Haplogroup R1a-L657
Y haplogroup can be determined from analyzing the Y chromosome of a male. Non recombining portion of Y chr is passed on only from father to son, and thus paternity can be determined. Once in many generations, one male undergoes one or many mutations in the Y chr (eg C -> T, G->A etc) and thus a subclade is born. This mutated subclade will now pass on through his son.
This is how R-L657 was born from Z93. (Brackets denote ISOGG notation)
R>R1>R1a>R1a1>R1a1a>R1a1a1>R1a1a1b>
> Z93(R1a1a1b2)> Z94(R1a1a1b2a)> Y3(R1a1a1b2a1)> L657(R1a1a1b2a1a)
Sample id I6942 is the only R-L657+ (+ means DERIVED or presence for the particular mutation, - means ANCESTRAL or absence of that mutation) found in ancient DNA so far. R-L657 is that subclade of R1a which is now common in Indian subcontinent and the middle east, but not found in modern Europe or ancient europe/steppe. The ISOGG name for it is R1a1a1b2a1a. Technically, I6942 is + for R-Y5 (R1a1a1b2a1a1a1~) and R-Y928 (R1a1a1b2a1a1a1f~) which implies that it is + for L657 as well.
It is safe to say that R-Y3 & R-L657 and its subclades were born in the Indian subcontinent given its absence in modern and ancient steppe and Europe (except Indian immigrants and the Romanis). I have always maintained that given that the formation and spread of R-Y3 is 2600 bce and R-L657 is 2200 bce respectively (YFull), that their paternal ancestors (Z93 and Z94) were somehow present in Indian subcontinent but without the presence of autosomal steppe ancestry which enters Indian subcontinent only post 1500 bce.
How did R-L657 reach India?
Here, it is worth noting that the oldest Z93 samples have been found in Fatyanovo culture, Russia dating to 2500 bce. So it is likely that this Z93 did enter India around the same time from Russia, however it would not be accompanied by any noticeable change in autosomal ancestry. Autosomal ancestry is that which recombines and is present in chr 1-22, and is passed 50-50% from both parents to child. Steppe autosomal ancestry only enters india post 1500 bce.
The other option of course is that India was the source of R1a to Europe, to prove which we need tonnes of ancient DNA which we don't have. Europe also has ancient samples of ancestors of Z93, like Z645 and M417 and so on, which makes european origin of R1a much more likely. So I will not consider this option yet, although its plausible. There can always be haplogroups which reach multiple places due to some travel savvy ancients

This has happened before, we see y hg J2 and J1 (and subclades) from Iran, caucasus and SC asia neolithic in european samples in Karelia_EHG and Austria LBK, Hungary_Sopot without any apparent autosomal ancestry from the source regions. Autosomal ancestry from a parent can dilute to ~0% easily in 7 generations if the sons end up marrying local women (50%>25%>12.5%>6.25%>3.125%>1.5%>~0) ie 150 years.
Some make the argument that the lack of R1a in Shahr Sokhta, BMAC, Turan eneolithic proves that R1a was absent in this region. To which my response is that these regions saw population turnover starting 4th mill bce or even earlier by a population like Tepe Hissar/Seh Gabi to the tune of >60% and a resulting Y HG turnover. This is the reason why west asian Y haplogroups like E1b and G2 appear in the record in the regions adjoining NW India. This turnover never happened in Indian subcontinent as Indians lack this anatolian heavy component. So I don't consider this evidence very strong, more data is needed.
SUMMARY
1. Sintashta component not correlated with R1a in RoopkundA samples, whereas Irula component is.
2. J2a/J2b significantly correlated with higher steppe ancestry in RoopkundA samples, steppe ancestry mediated through females.
3. I discuss possibilities regarding R-L657 reaching India.
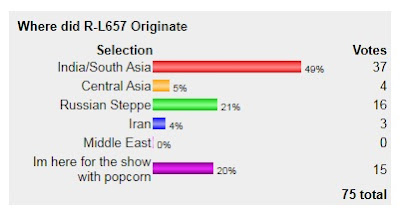
With this, I present the results of the L657 poll, I guess it's clear that Indian subcontinent won this one.
 Also read
Scratching my head! - Criticism of Narsimhan VM et al 2019 - Pt 1
The West/East Divide among Indo-Aryan languages
Also read
Scratching my head! - Criticism of Narsimhan VM et al 2019 - Pt 1
The West/East Divide among Indo-Aryan languages
1 Harney, É., Nayak, A., Patterson, N.
et al. Ancient DNA from the skeletons of Roopkund Lake reveals Mediterranean migrants in India.
Nat Commun 10, 3670 (2019).
https://doi.org/10.1038/s41467-019-11357-9
 archive.md
archive.md

