Continued from [] https://defenceforumindia.com/threa...ces-of-indus-valley-people.83063/post-2065110
PREMENDRA PRIYADARSHI.
La Genetique Scandale.
PART 2
Manipulation with data and Concoction
The Defective Design of Research: Systemic Bias
Bias in the modern scientific researches and concluding narratives is the rule rather than exception (Mullane and Williams, Eds., Biochemical Pharmacology, Elsevier Journal, 2013; Smith and Noble, British Medical Journal, 2014; Pannuki and Wilkins, Plastic and Reconstructive Surgery 2010). Narasimhans’ paper is the first class example of how biases can be operative at all the levels and stages of research. If all the manipulations and biases in the book is listed it will form a 1000 page volume, which is not my intention to do. Because will consume at least one year of my time writing all that. Some very obvious things, which even lay readers can understand will be noted below.
The Admixture Analysis and the Identification of Component
In the admixture Analysis presented by them, they have identified some primary components or populations genetically, which were to act as components for the formation of other populations by admixture later in the history. We shall first see only three of them :
Line 197 (BioRxive) . “Iranian agriculturalist-related”: represented by 8th millennium BCE pastoralists from the Zagros Mountains of Iran (17, 18)
Line 201 (ibid). “West Siberian Hunter-Gatherer (West_Siberian_HG)-related”: a newly documented deep source of Eurasian ancestry represented here by three samples
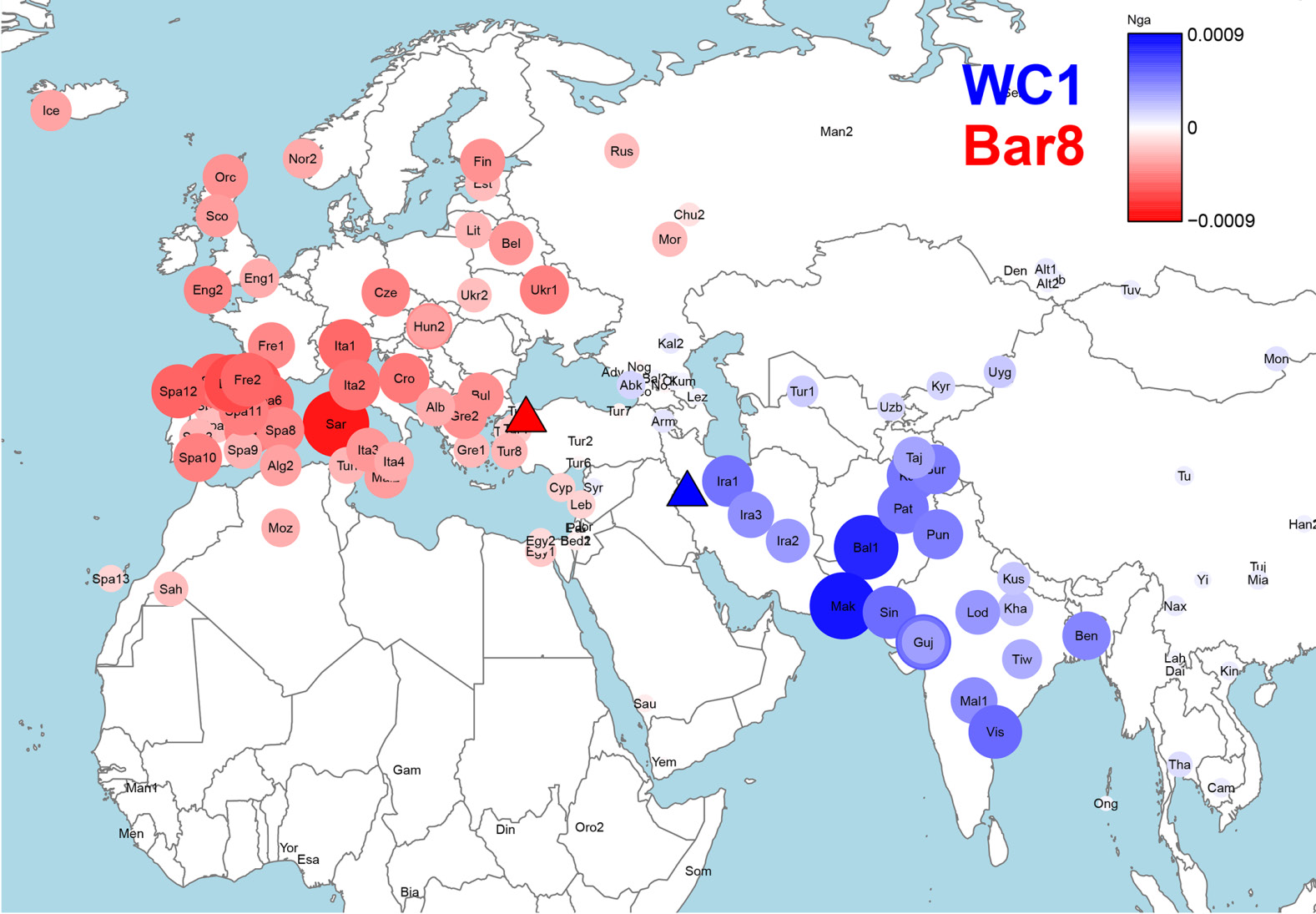
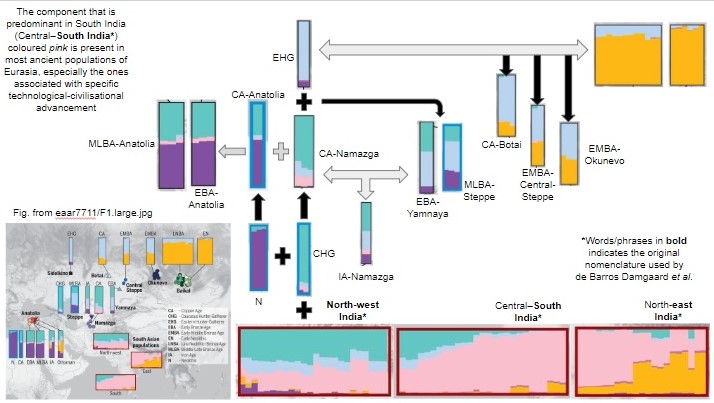
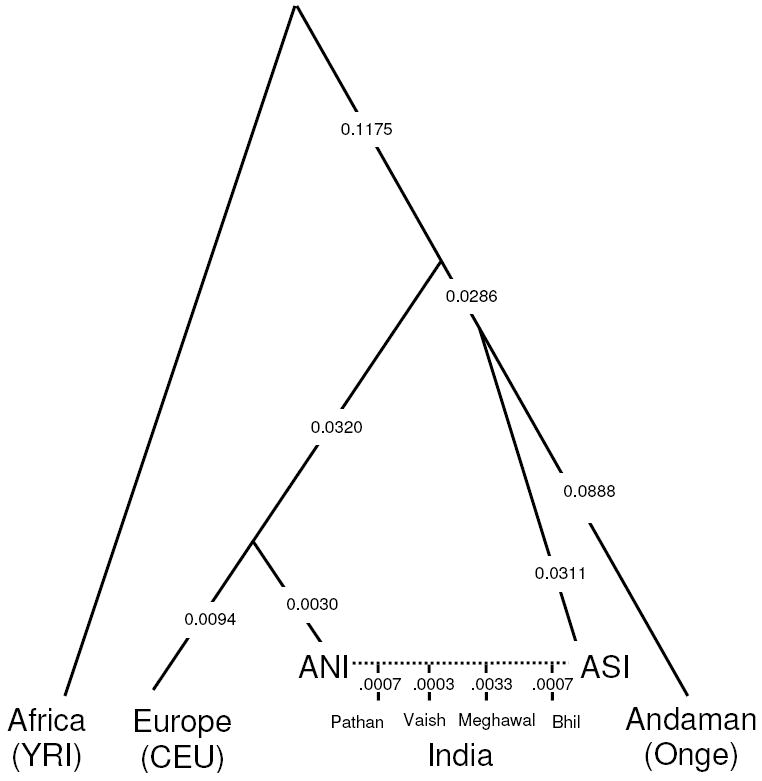
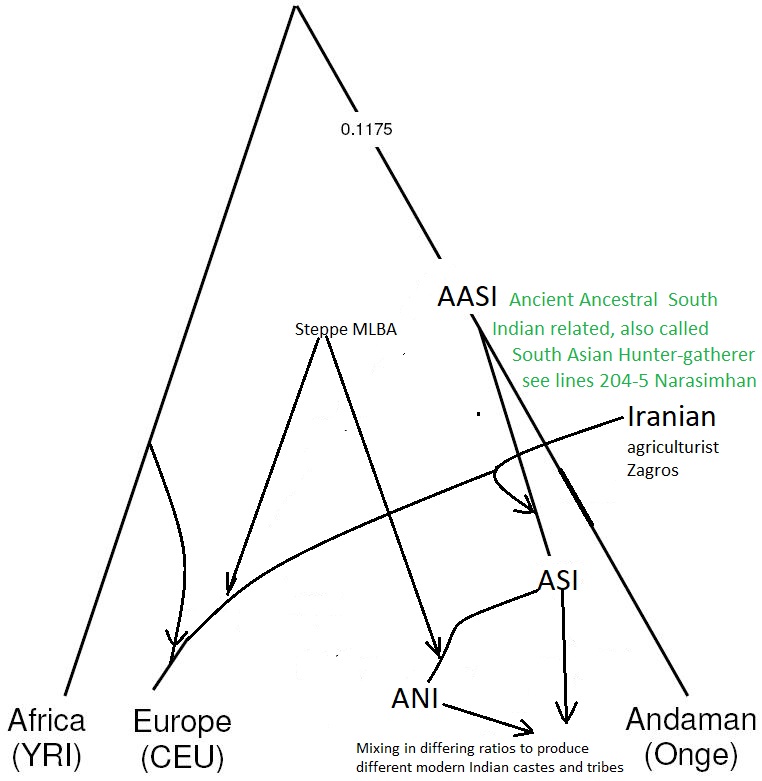
Line 204 (ibid) . “Ancient Ancestral South Indian (AASI)-related”: a hypothesized South Asian Hunter-Gatherer lineage related deeply to present-day indigenous Andaman Islanders (19) [The Narasimhans have used the DNA of the ONGE Tribe of the Andaman Islands to represent this gene pool.]
ONGE
The Narasimhans (a handy word in lieu of Narasimhan and colleagues) have identified Ancient Ancestral South Asian- (AASI)-related population by the modern Onge tribe’s genome, which live in the Andaman Islands today, and which are genetically related to the Papua New Guinea tribal peoples from the antiquity. Surprisingly the Narasimhans’ hypothesis that there were people having identical genomic constitution as the modern Onge living in India until the Iranians and the steppe people arrived and admixed with them. These Indians have been labelled as the AASI in the paper. This is a new invention (Neologism) uniquely invented for the occasion. These Onge people of North and South India admixed with Iranian and mid-to-late BA steppe genetic group to produce the modern Indian populations, Narasimhans conclude.
In their hypothesis the admixture of the Onge-genome with the Iranian one gave rise to the South Indian (Dravidian) population; and the admixture of this with the Middle-to-late-Bronze Age steppe population gave rise to the North Indian (Aryan) population, they hypothesize. In other words it is the resurrection of Elamite theory of the origin of Dravidian in addition to the Aryan Invasion Theory.
However the real story is different. Onge people could have been of the same genetic composition as the Indians about 70,000 to 50,000 years back, but not today, not even 10,000 years before today. These people (Onge) are the remnants of the people who migrated to the Anadaman Islands, and also to the southeast Asia about 70,000 to 50,000 years back. Some had stayed in India; some had arrived into Andaman (ancestors of the Onge and Jorwa); and some others would migrate eastward towards Papua New Guinea and Australia. But they did not stop there. They migrated further east to South America.
Thus the Onge separated from the rest of the humanity about 50,000 to 70,000 years ago, noted Thangaraj et al (2005) in their genetic study: This was the same time when the Australians (and the Papuans) separated from the rest of the humanity. They are considered Australasian family, and must be considered much more remotely connected to the Ancient Ancestral South Asians than the European Cro-Magnons, Siberians, Iranians and the Central Asians who branched off from the main basal Eurasian trunk much later than the Onge did.
It is useful here to recollect what Thangaraj wrote about the Onge: “Our data indicate that two ancient maternal lineages, M31 and M32 in the Onge and the Great Andamanese, have evolved in the Andaman Islands independently from other South and Southeast Asian populations. These lineages have likely been isolated since the initial penetration of the northern coastal areas of the Indian Ocean by anatomically modern humans, in their out-of-Africa migration –50 to 70 thousand years ago.” [Thangaraj et al, 2005, Reconstructing the Origin of Andaman Islanders, Science, 308 (5724): 996.]
Thangaraj also noted that over the period of time the genetic composition of the populations of the Andaman-Onge and Indian-Mainland population drifted from each other due to changes taking place in the two independently. They note: “Analysis of the complete mtDNA sequences shows that none of the coding region mutations defining these two haplogroups overlap with the known Indian or East Asian mtDNA haplogroups (1–5). In our survey of 6500 mtDNA sequences from mainland India, none of the M lineages carried the coding region mutations specific to M31 and M32 (6).” (Thangaraj 2005). Thus we can see that genetically the Onge had deviated a lot from mainland India over the last 50,000 years of the separation. The Onge people are short-stature and dark skinned. (Thangaraj is one of the authors in Narasimhan’s article too).
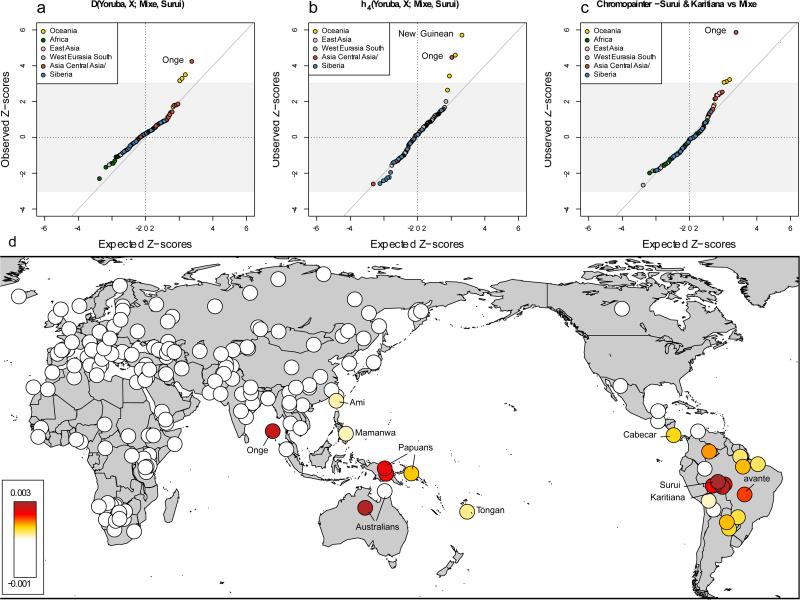
In fact the article which claimed that the Onge had migrated east even up to South America had been authored by a group of authors which included the doyen David Reich himself.
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC4982469/figure/F1/
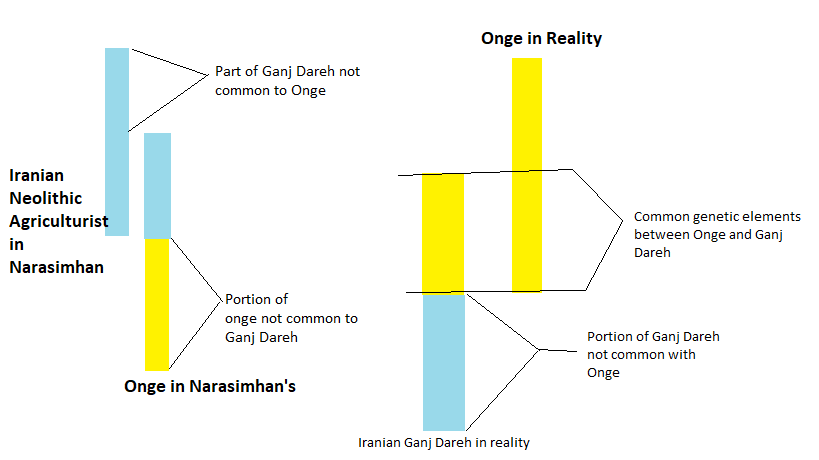
Thus considering Onge as the Indian population at the onset of Neolithic marred the total work, and vitiated all the results. Instead of considering the Onge as single component, their admixture analysis found that it is an admixture of Iranian DNA. This error occurred because the authors failed to recognise that the Onge and the Iranians both had split from the mainland Indians, although in different eras. Hence there must be some elements common in the three the Iranian, the Mainland Indian and the Onge populations. However there would be some portions which would be distinct in the three populations.
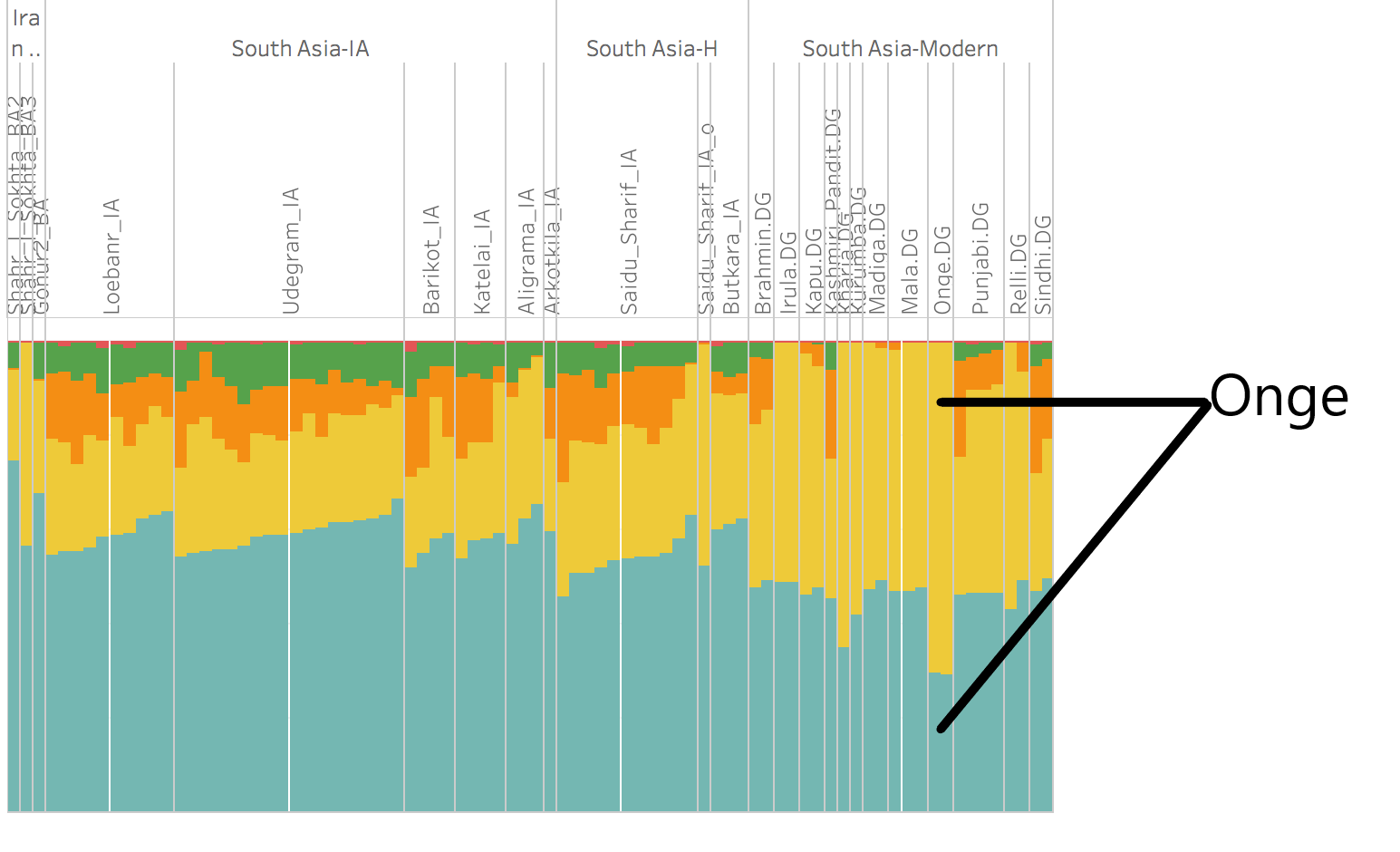
Figure: Onge as shown in the Admixture Analysis in the Figure S3.37; page 166 of the Supplement of the Narasimhan et al. Onge highlighted and pointed out by us. In this figure, Onge has almost the same components as the Mala, Irula, Shaidu-Sharif (Iron Age_0) and Shahar-i-Sokhta BA3, except that the latter ones have greater proportions of the Iranian component.
This failure to appreciate some commonality (due to origin) between Indian and Iranian DNAs resulted in considering the Iranian farmers as unmixed pure component. It was also compounded by the false and erroneous belief that nobody could ever have gone out of India. In an effort at negating the Indian components in the Iranian Agriculturists the authors used the parameters in such way as that it read Iranian Agriculturists made up of a single component, coloured teal (light blue) in the Admixture Analysis.
Such distortion in the calibration vitiated the whole result in such a way that the mono-component genomes of the Onge (Andamanese) started showing made up of two components. One of them was blue (Iranian Agriculturists) as depicted in the histogram above.
How it happened can be seen in the figure below:
Figure: Onge (Andaman) as depicted in Admixture Analysis by Narasimhan, and as in reality it should have been. Commonality between Ganj Dareh (Iranian Agricult.) and Onge (Andaman) only means that a large proportion of the gene in the Ganj Dareh was same as Ancient Ancestral South Asians, today represented in mainland India by Mala, Irula etc, and in the Andaman as Onge. The Ancient Iranian Agriculturists have been shown as light blue (teal) colour and the Andaman-Onge are half light-blue (teal) and half yellow.
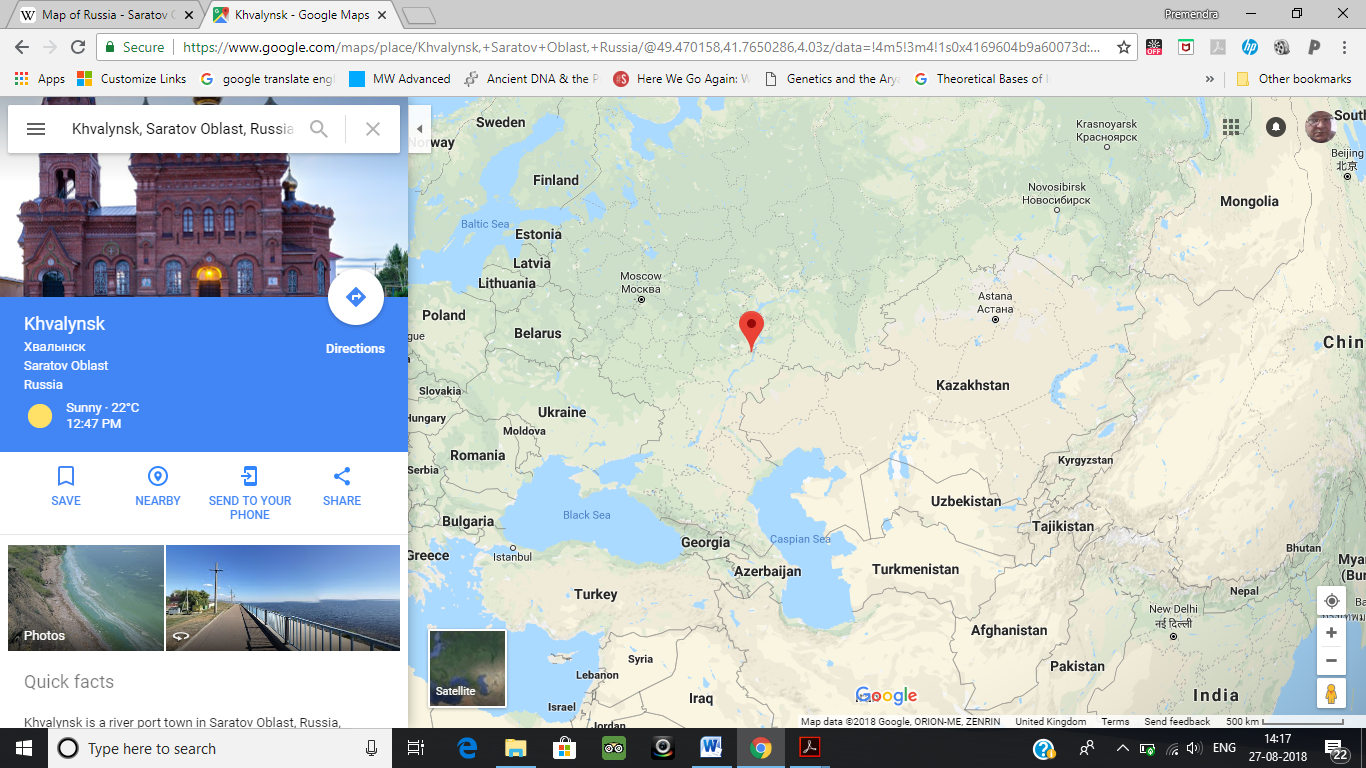
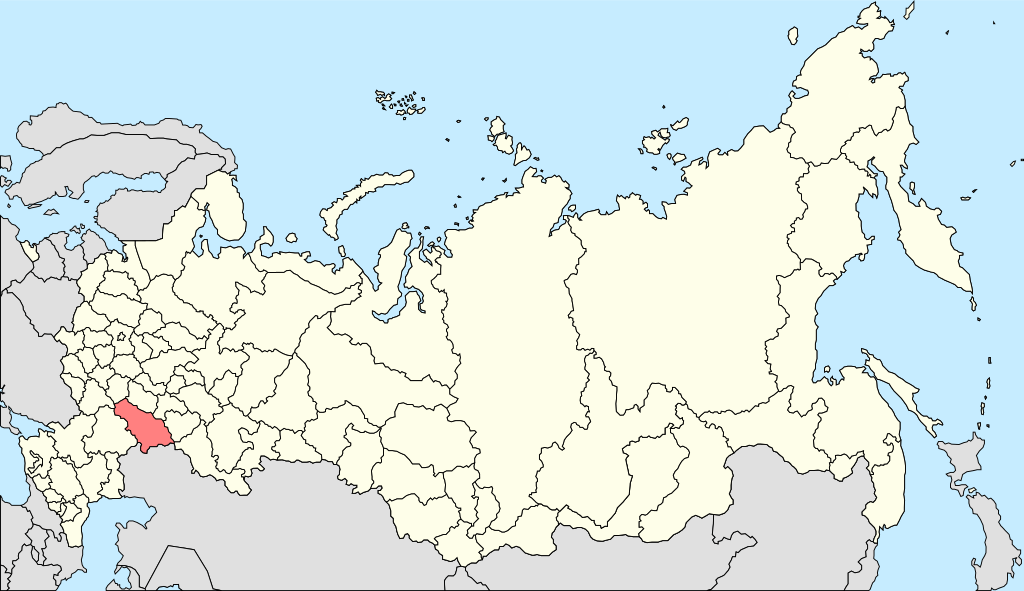
Iranian (Jagros Neolithic): Ganj Dareh
The Ganj Dareh has been postulated as the one of the principal components in the Narasimhan study. However it has clear ancestry from Northwest Indian sub-continent, where Pakistan is located today. Narasimhan’s picture itself shows that the Ganj Dareh is only a sub-set of the larger set which is Onge, Mala etc Indian samples.
Unfortunately for the Narasimhans, the Ganj Dareh (Iran Neolithic) DNA was studied by another group of workers whose work was published in the Nature (which is not a biased journal like Science) found that:
“The mitochondrion of GD13a (91.74X) was assigned to haplogroup X, most likely to the subhaplogroup X2, which has been associated with an early expansion from the Near East and has been found in early Neolithic samples from Anatolia, Hungary and Germany.” (Llorente 2016)
“GD13a did not cluster with any other early Neolithic individual from Eurasia in any of the analyses.” (Llorente 2016)
[Interpretation: We need to remember that no South Asian Neolithic DNA has been reported so far. Thus if GD13a a matched none, possibly only one which remains to be tried is the South Asian Neolithic DNA.]
“We further investigated the relationship between GD13a and Caucasus Hunter-Gatherers using D-statistics to test whether they formed a clade to the exclusion of other ancient and modern samples (Table S4). A large number of Western Eurasian samples (both modern and ancient) showed significant excess genetic affinity to the Caucasus Hunter-Gatherers, whilst none did with GD13a. Overall, these results point to GD13a having little direct genetic input into later European populations compared to its northern neighbours.” (Llorente 2016)
[Interpretation: Thus the oldest Iranian Agriculturists of the Ganj Dareh had not directly migrated to Europe, although their mitochondrial DNA X2 certainly contributed to the modern European population. Other studies have shown (see below) that the Iranian Neolithic DNAs had migrated to Levant and Anatolia, and the Caucasus as well as the steppe. Clearly the Ganj Dareh DNAs were not the gross representative of the Iranian Neolithic, but they represent a segment which was very small and did not make much impact on later Europe, Caucasus or the steppe. It might have represented just a small number of the emigrants from a single village in Baluchistan or other part of the northwest South Asia, and might have represented only a tiny fraction of genetic variation which South Asia had at that time. On the other hand the people reaching the other locations in Zagros (Iran) at Neolithic might have originate from other villages of Pakistan/ Afghanistan resulting in the difference in the genetic composition within the Zagros Neolithic populations.]
“The individual analysed here was part of burial 13, which contained three individuals, and was recovered in level C in 1971 from the floor of a brick-walled structure. The individual sampled, 13A (referred to as GD13a throughout the text), was a 30–50 year old female; the other individuals in the burial unit were a second adult (13B) and an adolescent (13). The site has been directly dated to 9650–9950 cal BP, and shows intense occupation over two to three centuries. The economy of the population was that of pastoralists with an emphasis on goat herding7. Archaeobotanical evidence is limited but the evidence present is for two-row barley with no evidence for wheat, rye or other domesticates. This implies that the overall economy was at a much earlier stage in the development of cereal agriculture than that found in the Levant, Anatolia and Northern Mesopotamian basin.” (Llorente 2016)
[Interpretation: This information refutes the claim by the Narasimhans that the Mehrgarh farming culture had been borrowed from Anatolia (Turkey) through Iran (Ganj Dareh). The date cited above gives a date of 7,850 BC (mean). It may be noted that the Mehrgarh oldest layer has a date of 8,707 BC (mean).
While the Ganj Dareh Iranian people had only two-row barley (see above) at 7850 BC, the Mehrgarh had six-row barley at 8700 BC, which is an advanced stage of agricultural development and domestication of barley (Upinder Singh 120; Jarrige 2008). Jarrige wrote citing Lorenzo Costantini:
“Lorenzo Costantini has shown that the plant assemblage of Period I is dominated by naked six-row barley which accounts for more than 90% of the so far recorded seeds and imprints. He has also pointed out the sphaerococcoid form of the naked-barley grains with a short compact spike with shortened internodes and small rounded seeds.
According to him, such characteristics in the aceramic Neolithic levels can be ascribed to probably cultivated but perhaps not fully domesticated plants. Domestic hulled six-row barley (H. vulgare, subsp. vulgare) and wild and domestic hulled two-row barley (H. vulgare subsp. spontaneum and H. vulgare subsp. distichum) have also been recorded, but in much smaller quantities. According to Zohary quoted by R.H. Meadow, the distribution of wild barley extends today to the head of the Bolan Pass. It is therefore likely that local wild barleys could have been brought under cultivation in the Mehrgarh area. Costantini has also identified a small amount of domestic einkorn (hulled: Triticum monococcum), domestic emmer (hulled: T. turgidum subsp. dicoccum) and a free-threshing form which can be referred to as Triticum durum (Fig. 10). ” (Jarrige 2006)
Thus these people of Iran had arrived here about 1000 years after the Mehrgarh took off. The Ganj Dareh site had been occupied by only a short period of 100 to 300 years (mean 200 years). On the other hand the Mehrgarh shows a continuous occupation till late. It may be noted that the domestication of the goat is not possible to take place in 200 or 300 years and about a thousand years are required to give domestication features to the animal skeletons. Clearly the people of Ganj Dareh were not local, and had arrived from somewhere else.]
“ADMIXTURE and outgroup f3 statistics identified Caucasus Hunter-Gatherers of Western Georgia, just north of the Zagros mountains, as the group genetically most similar to GD13a (Fig. 1B,C), whilst PCA also revealed some affinity with modern Central South Asian populations such as Balochi, Makrani and Brahui (Fig. 1A and Fig. S4).” (Llorente 2016)
[It is possible to interpret it as the Ganj Darh coming from a region within the locations Brahui, Baluchistan and Makaran of South Asia, now in Pakistan. Mehrgarh was in Baluchistan.]
“Also genetically close to GD13a were ancient samples from Steppe populations (Yamanya & Afanasievo) that were part of one or more Bronze age migrations into Europe, as well as early Bronze age cultures in that continent (Corded Ware) in line with previous relationships observed for the Caucasus Hunter-Gatherers.”
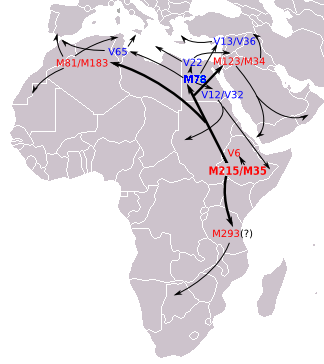
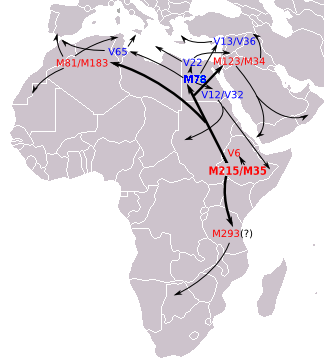
[Yamnaya and Afanasievo of steppe/ Central Asia were much later than Ganj Dareh (Iran). The resemblance could be due to either the Iranian early farmers migrating into the steppe. But these people could have developed from the migrations from the northwest India/ Afghanistan/ Pamir region. The Pamir is a likely source of R1b which migrated by northern Iranian/ Turkmenistan wet corridor to south of Caspian and then from there to Armenia, Anatolia and then Southern, Western and Central Europe. Yamnaya and Afanasievo belong to R1b Y-DNA. See figures below.]
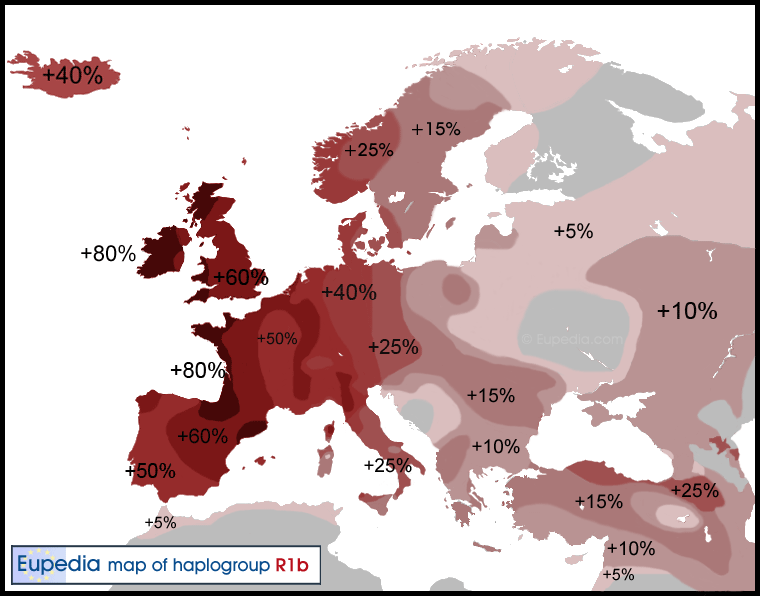
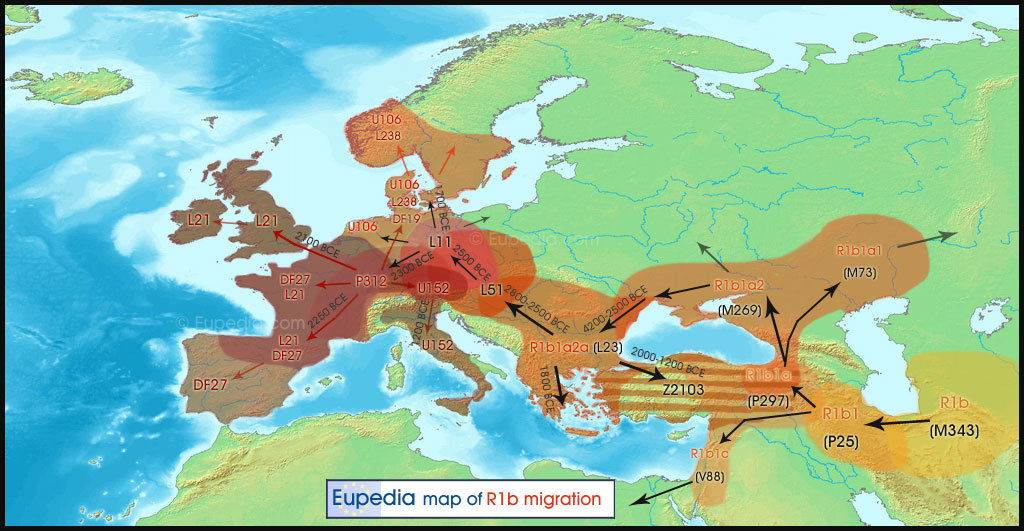
Figure: Eupedia map of R1b distribution in Europe. It is dominant in those parts of Europe which speak Centum languages like Scottish, Irish, Welsh, Italic (French, Spanish, Portugese, Italian), Greek, Anatolian, Armenian and Germanic (English, Norwegian, German etc). The steppe language Tocharian was also Centum which was spoken by the Yamnaya and the Afanasievo people in the Bronze Age.
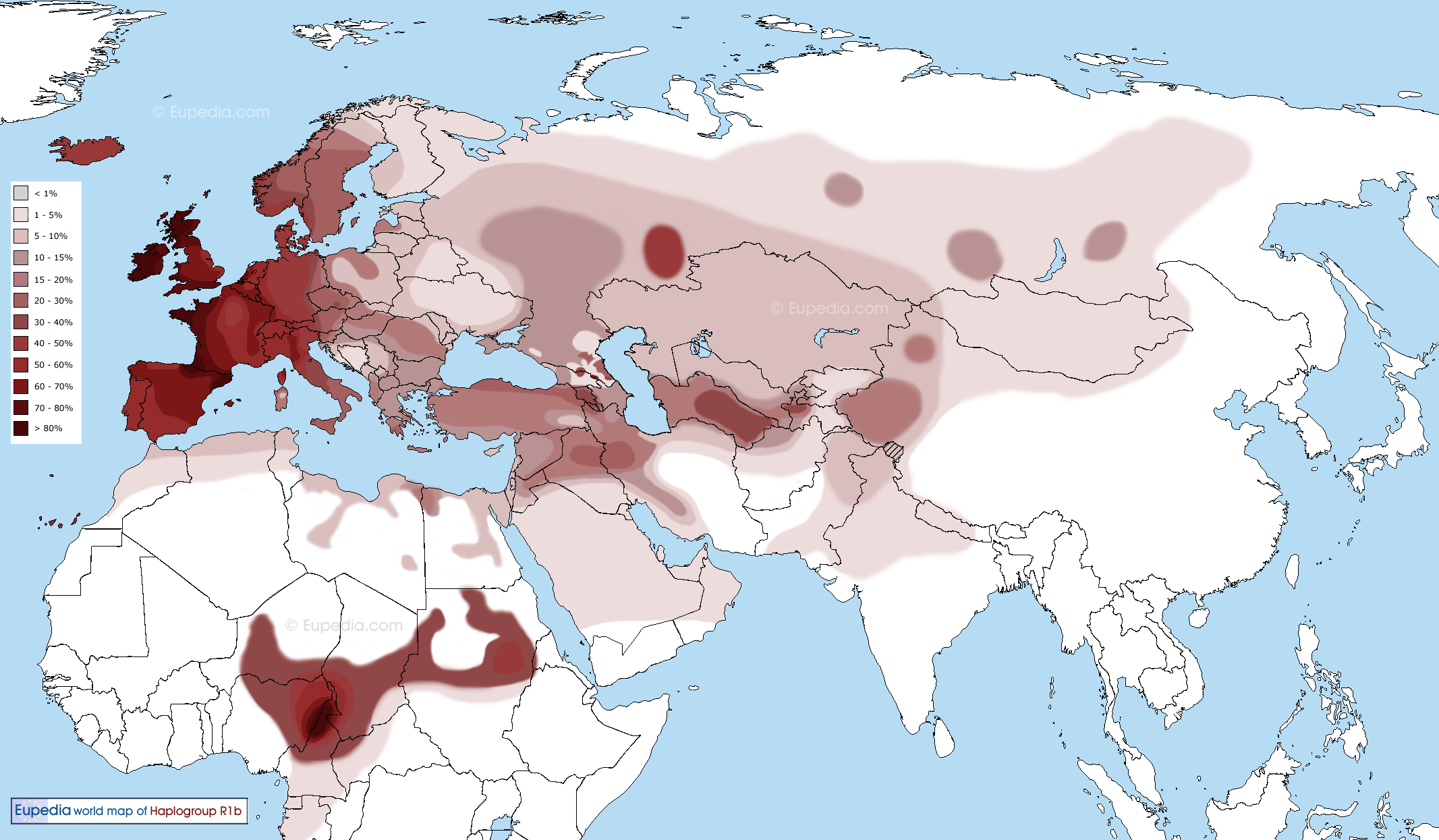
Figure: source Eupedia. The R1b might have originated in the region to the north of Kashmir i.e. Tajikistan, and migrated through Tajikistan to Armenia and further. It also went to Central Africa. In steppe and Central Asia it was replaced by later arrivals in Late Bronze Age by the R1a, which is dominant only in the Satem speaking groups like Russian, Ukrainian, other Slavic, Latvian, Lithuanian, Polish, Iranian, Indic etc.
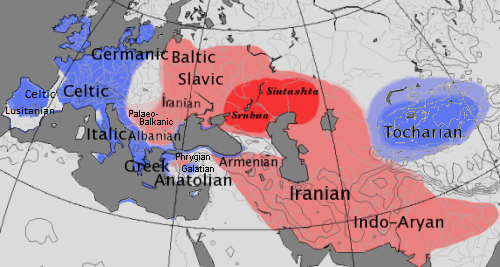
Figure: Source Wikipedia. Map showing distribution of Centum and Satem branches of Indo-European
Figure: R1b Migration map suggested by Eupedia
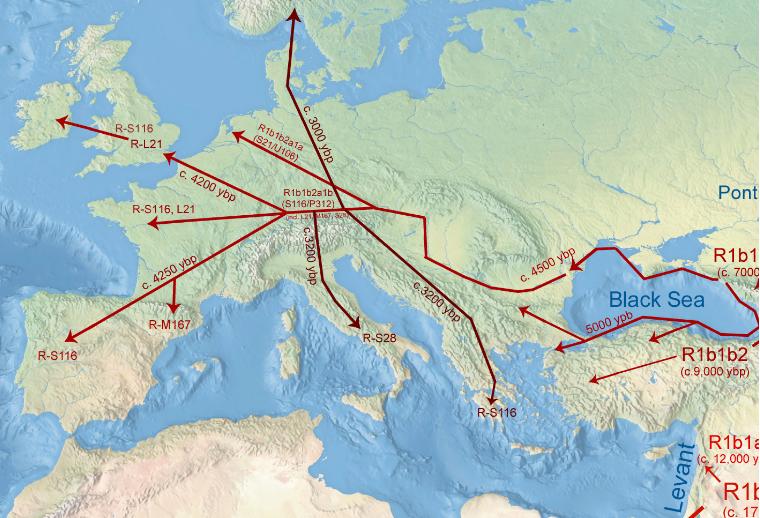
Figure: R1b Migration map as suggested by Wiki
Now we can also see regarding Ganj Darh what Llorente had to say further:
“We further investigated the relationship between GD13a and Caucasus Hunter-Gatherers using D-statistics to test whether they formed a clade to the exclusion of other ancient and modern samples (Table S4). A large number of Western Eurasian samples (both modern and ancient) showed significant excess genetic affinity to the Caucasus Hunter-Gatherers, whilst none did with GD13a. Overall, these results point to GD13a having little direct genetic input into later European populations compared to its northern neighbours.” (Llorente 2016)
[Later Europeans are products of another wave of migration namely R1b which came later in Bronze Age.]
“Thus, GD13a is the descendant of a group that had relatively stable demography and
did not suffer the bottlenecks that affected more northern populations.” (Llorente 2016) [Interpretation: This line points to India.]
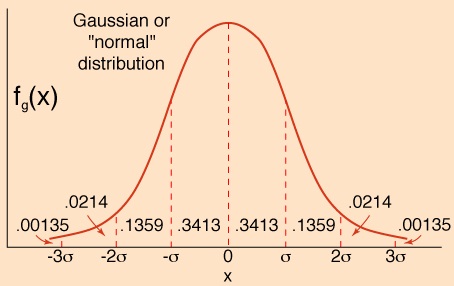
Figure: It may be understood from the Gaussian Normal Distribution curve that it is quite normal finding in any normal data that the extremes look different. It does not mean that they belong to two populations. However the naïve would take them as two populations.
What about other characters of the Ganj Dareh lady? Llorante noted the skin colour of the Ganj Dareh lady. :
“She lacked the derived variant (rs16891982) of the SLC45A2 gene associated with light skin pigmentation but likely had at least one copy of the derived SLC24A5 allele (rs1426654) associated with the same trait. The derived SLC24A5 variant has been found in both Neolithic farmer and Caucasus Hunter-Gatherer groups5,15,24 suggesting that it was already at appreciable frequency before these populations diverged. Finally, she did not have the most common European variant of the LCT gene (rs4988235) associated with the ability to digest raw milk, consistent with the later emergence of this adaptation5,15,21.” (Llorante 2016).
Clearly she had the light skin colour gene SLC24A5 allele which produces light skin colour in the Europeans and the Indians (Basu Mallik et al 2013). This gene was not found in the Europeans until late Bronze Age. It was not present in the La Branda human of 5000 BC. However it was found present in many European people between 3000 BC and 1000 BC (Allentoft). This means the Ganj Dareh were not ancestral to the early Neolithic people of north of Black Sea who entered East Europe replacing the hunter-gatherers at about 5000 BC. In my hypothesis the light skin colour gene SLC24A5 originated in South India long back, and it migrated to other places including even Ethiopia from India.
https://en.wikipedia.org/wiki/SLC24A5
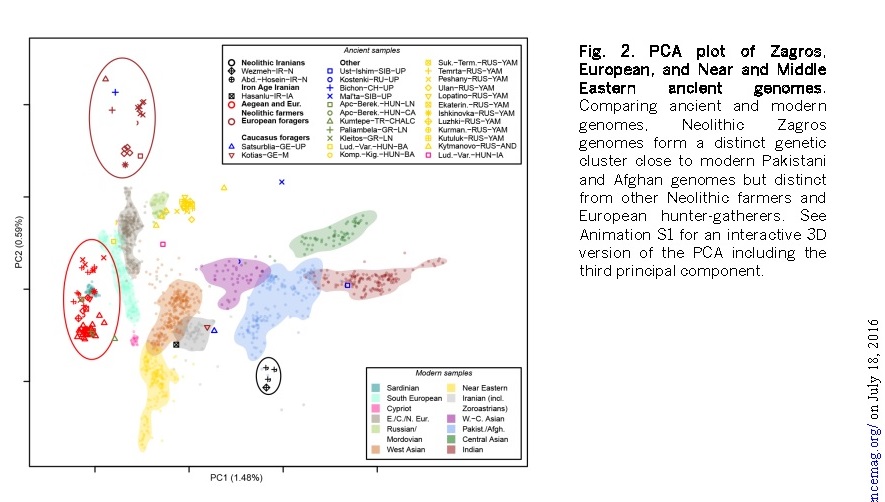
Another worker Broushaki (2016) noted that the Iranian Neolithic people from Wezmeh Cave were related to the Pakistani and Afghan people particularly to the Zorastrians of Iran origin now living in India. “These people are estimated to have separated from Early Neolithic farmers in Anatolia some 46-77,000 years ago and show affinities to modern day Pakistani and Afghan populations, but particularly to Iranian Zoroastrians.” Clearly the Zagros (Iran) farmers had not arrived from Anatolian farmer community of the Anatolia Neolithic. In fact they are deeply related to Indian population.
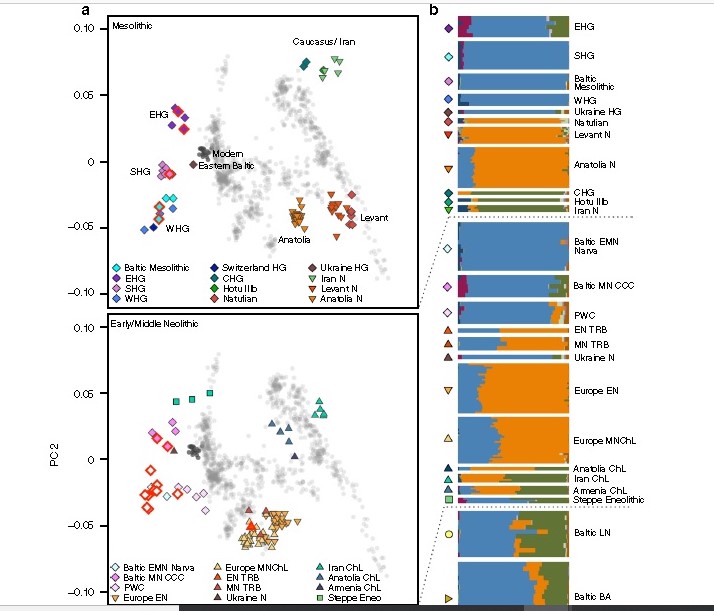
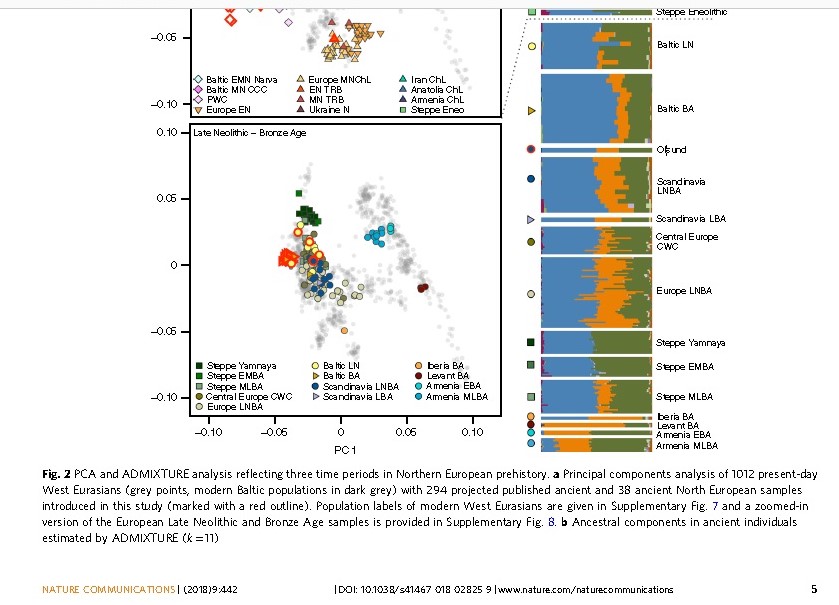
Figure Neolithic Iran as compared to Indian genome by Broushaki 2016
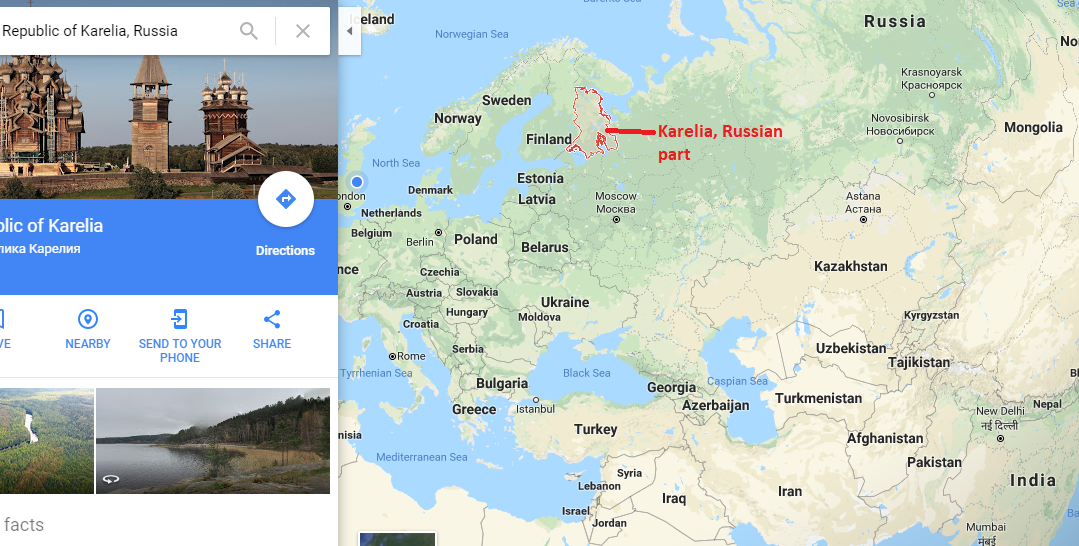
The Western Siberian Hunter Gatherers
Narasimhans have selected the following samples as representing the West Siberian DNA:
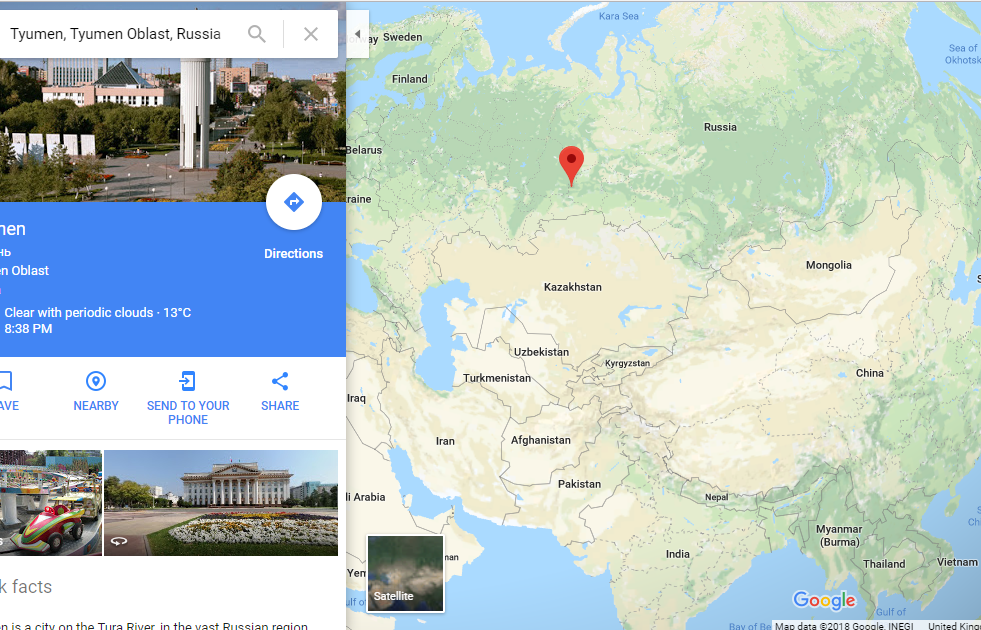
Sosnoviy-Ostrov, western Siberia, Russia (n=1); Tomsk10 (I5766): Date of 4230-3983 cal BCE (5261±33 BP, OxA-33486, d15N=+12.8 permil possible marine influence). Genetically female.; Tyumen Oblast, western Siberia, Russia (n=2) Tyumen1, Kurgan 1 (I1958): Date of 4723-4558 cal BCE (5805±25 BP, PSUAMS-2359), Genetically female; Tyumen50, Kurgan 6 (I1960): Date of 6361-6071 cal BCE [6335-6071 cal BCE (7330±40 BP, Poz-82198), 6361-6086 cal BCE (7355±40 BP, OxA-33489, d15N=+15.3 permil possible marine influence)]. Genetically female.
Location of Tyumen Oblast, the Ancient West Siberia HG genes
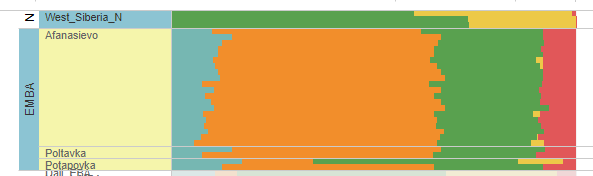
These places Sosnoviy- Ostrov and Tyumen Oblast by foot are about 3400 kms to the north of Kabul. They had yellow colour (AASI, Indian, Onge) component about one third quantitatively in the Admixture Analysis. To mislead people denial of this fact was done, not by changing the colour of the component, but by considering it an entirely different component although it looked same as Indian.
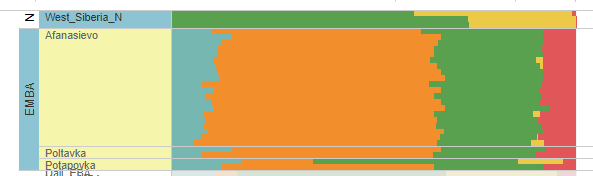
Figure: West Siberia admixture Analysis at the top. Source Narasimhan.
Apart from this the Barros Damgard too have provided another set of histograms for the Admixture Analysis of the same populations, with locations marked. This is more honest and correct and not tampered with.
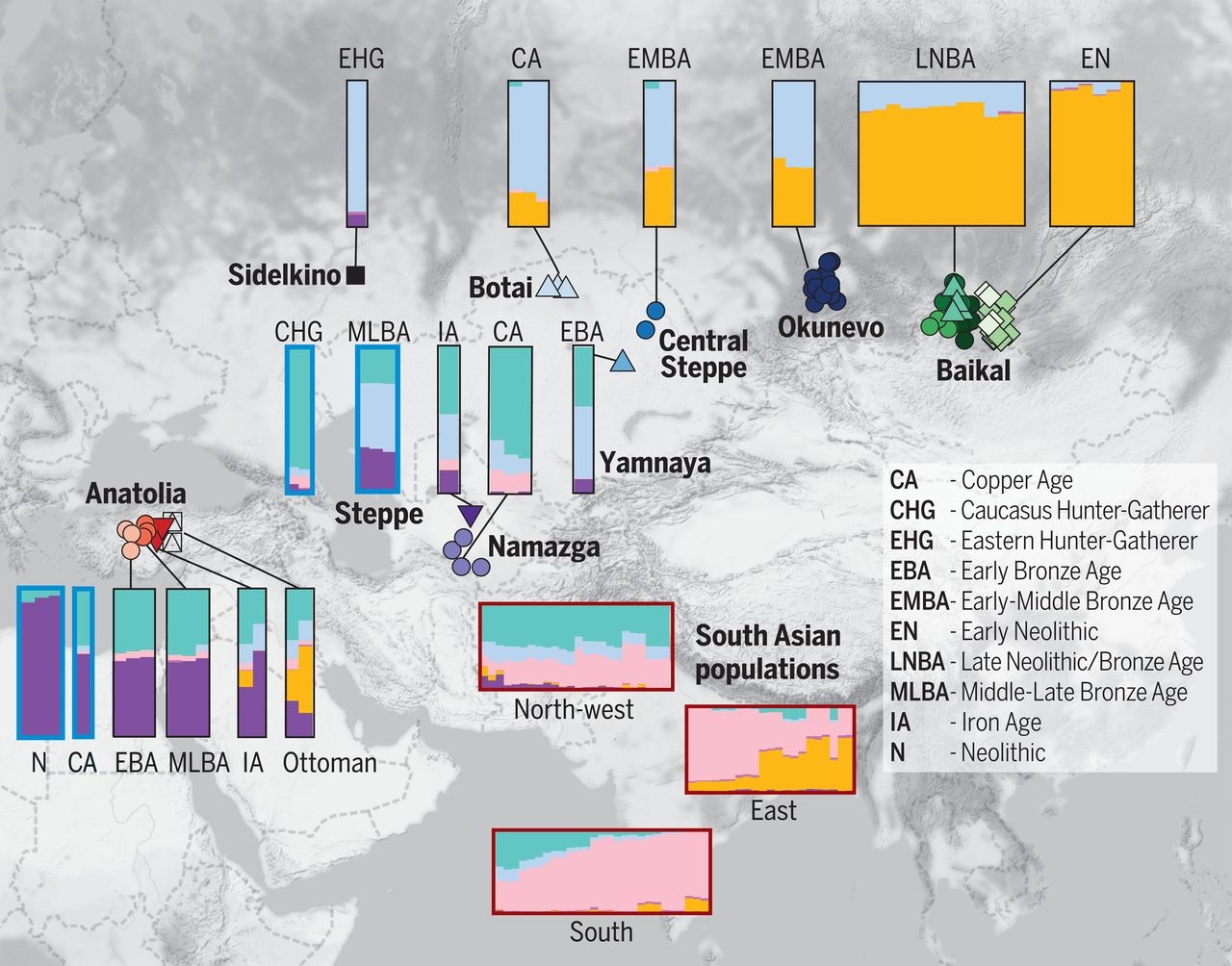
Figure: Barros Damgaard 2018, Admixture Analysis, Science
If rearranged, this picture gives the following outcomes:
This figure indicates that the Indian cline should be defined as East to South to Northwest in a folded shape or V shape. There is a gradual change in proportions of the golden, pink and teal (bluish-green) colours. Such arrangement indicates natural settlements with genetic movements not by migration but by the Brownian Movements of the genes.
If any arrival takes place, there is a breach in the cline, in the same way as Broushaki got one between Zagros and Anatolia during the Neolithic.
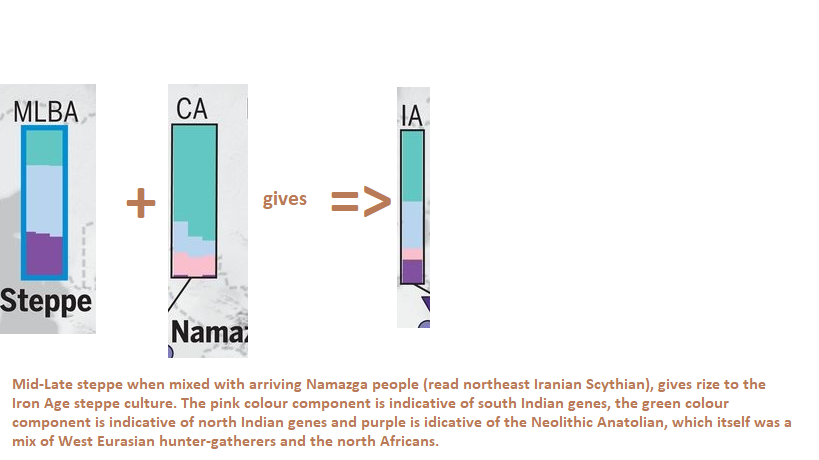
The further summations of the components indicates that the steppe may have originated from northwest India:
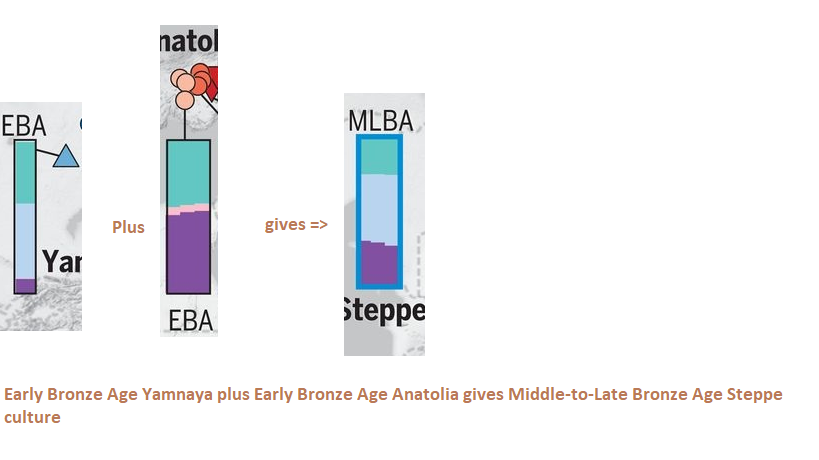
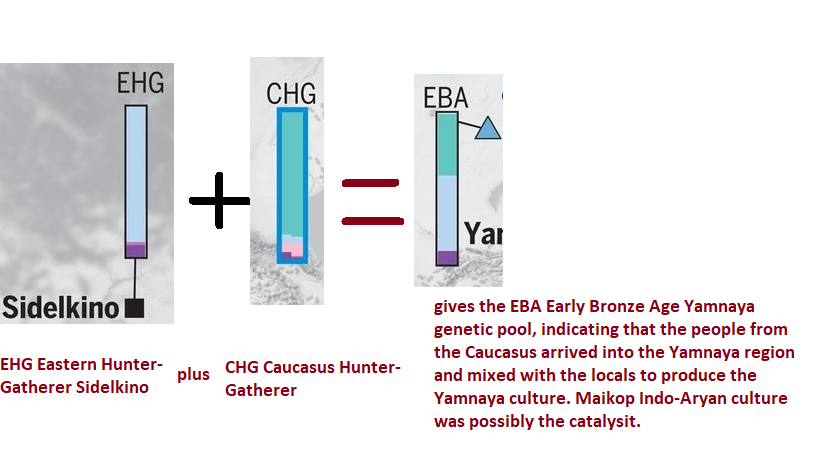
Figure: Eastern Hunter Gatherer (Sidel’kino location near Samara east of Volga river) plus Caucasus hunter gatherer gives if averaged, the Early Bronze Yamnaya. Clearly people coming through the Caucasus admixed with the local EHG to produce the Yamnaya culture. This happened during the R1b-Y-DNA expansion. Because the Yamnaya is mainly R1b. After reaching southern Caspian coast, the R1b people turned north. Established the Armenia Indo-European (Centum) and moved into Caucasus forming the Maikop (Maykop) culture in north Caucasus. It is believed that the Maikop people gave rise to the Yamnaya.
Figure: Admixture Analysis, Early Bronze Age Yamnaya plus Early Bronze Age Anatolia averaged gives the Middle to Late Steppe population.
Figure: MLBA steppe and Namazga Copper Age when averaged gives the Iron Age steppe culture.
We know from other studies that lot of the ancient samples of Y-DNA H1 lineage, which is typically Indian and most probably of South Indian origin later expanding to North India, have been found from Eneolithic to Bronze Age periods from locations in Anatolia, Middle East (e.g. Namazga), and North of Mongolia (Lake Baikal region, Shamanka). Clearly Indians had been migrating to wider regions of Asia much before the steppe culture took off during the Bronze Age period. (See Supplementary matter Excel Table aar7711_Table 14, of Barros Damgaard, Science 2018).
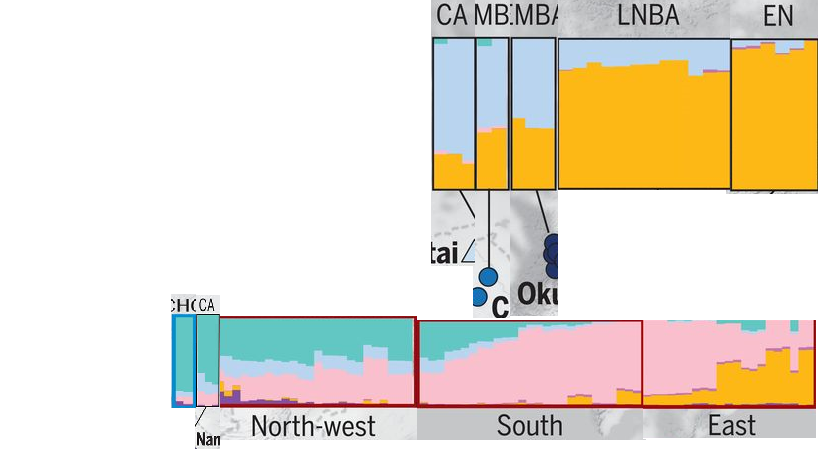
Thus we can conclude as this picture:
Figure: Barrow and Damgaard data rationally reorganised to produce understandable conclusions; (courtesy significant contribution from Dr Murali V.)
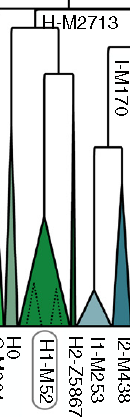
The H1 had a sibling H2 which has been found from Neolithic of Levant and Anatolia, and Sardinia. It has also been found from West Lake Baikal Shamanka region from Eneolithic period (sample number DA339 in Barros Damgaard 2018, se table). H3 is another branch which is found in the Romany. The early branch H0 which had split the earliest from the main trunk of H is also found in India only.
Family tree of Y-DNA H: Poznik 2016 Figure 2 enlarged